Archival Notice
This is an archive page that is no longer being updated. It may contain outdated information and links may no longer function as originally intended.
Home | Glossary | Resources | Help | Contact Us | Course Map
As previously mentioned, there are two analysis modes (classic Macintosh and advanced Windows NT modes). The differences between these modes are found in the sizing method and the flexibility of peak sensitivity settings.
Specifically the mode differences are:
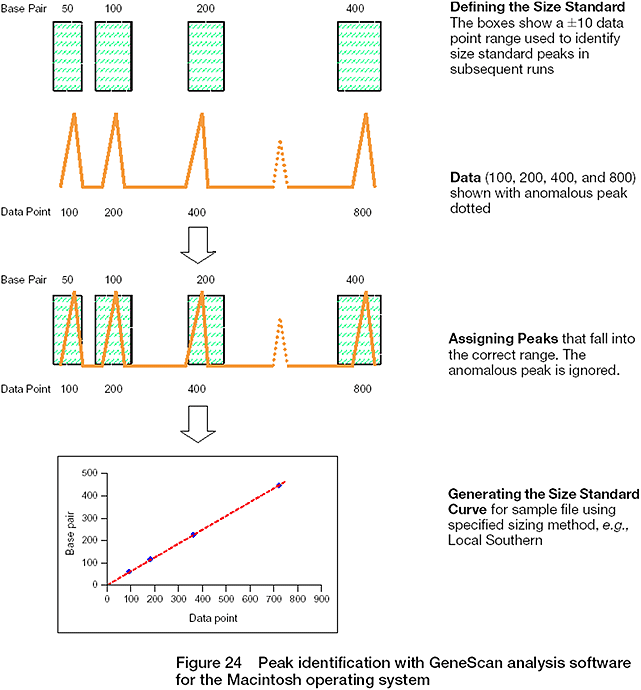
- In the classic mode, size calling is performed by matching the actual size standard fragments of the sample with a defined size standard that must be accurately labeled; it utilizes scan number to assign sizes.
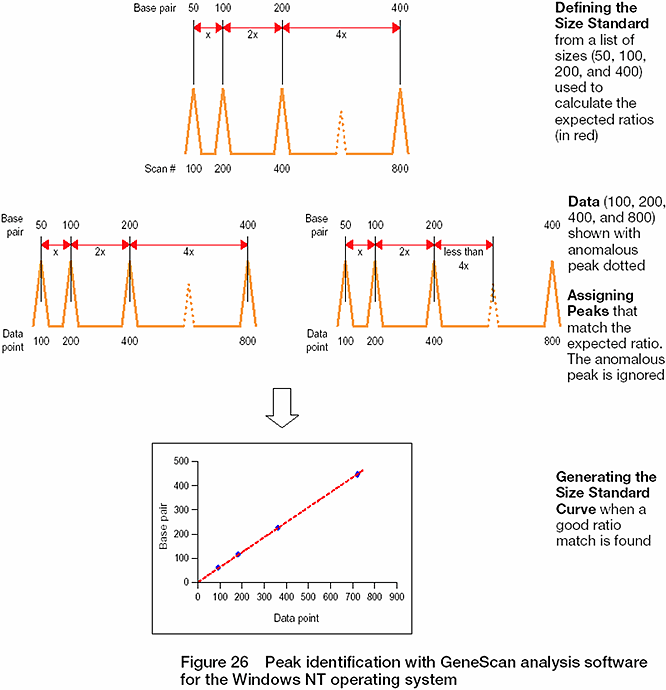
- In the advanced mode, size calling is performed using a function known as "ratio matching." Ratio matching uses an algorithm to determine the distance between the size fragments based on the entry of a set of size fragment values, where it uses the relative distance between the neighboring peaks to size.
The following excerpts from the Applied Biosystems User Bulletin on Size Parameters explain these differences.05
There are similarities between GMID and GS/GT software. However, GMID offers additional features, added flexibility, and efficiency through the combining of the programs. The expert system potential, inherent in the software, will continue to develop as new versions are released.
Note: |
|---|
| To become familiar with the use of GMID 3.2, it is recommended that analysts read through the User's Manual for v3.1 and User Bulletin for v3.2; focusing particular attention on the verification process and the software features and functions.07 |
Additional Online Courses
- What Every First Responding Officer Should Know About DNA Evidence
- Collecting DNA Evidence at Property Crime Scenes
- DNA – A Prosecutor’s Practice Notebook
- Crime Scene and DNA Basics
- Laboratory Safety Programs
- DNA Amplification
- Population Genetics and Statistics
- Non-STR DNA Markers: SNPs, Y-STRs, LCN and mtDNA
- Firearms Examiner Training
- Forensic DNA Education for Law Enforcement Decisionmakers
- What Every Investigator and Evidence Technician Should Know About DNA Evidence
- Principles of Forensic DNA for Officers of the Court
- Law 101: Legal Guide for the Forensic Expert
- Laboratory Orientation and Testing of Body Fluids and Tissues
- DNA Extraction and Quantitation
- STR Data Analysis and Interpretation
- Communication Skills, Report Writing, and Courtroom Testimony
- Español for Law Enforcement
- Amplified DNA Product Separation for Forensic Analysts