Archival Notice
This is an archive page that is no longer being updated. It may contain outdated information and links may no longer function as originally intended.
Home | Glossary | Resources | Help | Contact Us | Course Map
Mitochondrial DNA (mtDNA)
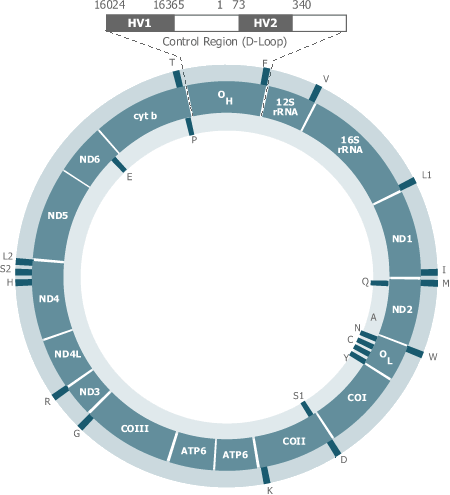
Typically, mtDNA is used to type samples such as teeth, bones, and hair that are normally intractable to standard STR (Short Tandem Repeats) analysis. The hypervariable control region of mtDNA used for typing contains a large number of linked SNPs (Single Nucleotide Polymorphisms) that are simultaneously determined by standard Sanger sequencing.
The coding region also possesses SNPs that can be useful in differentiating between samples that otherwise exhibit the most common Caucasian control region mtDNA haplotype. These coding region SNPs are less dense than in the control region and are typically typed using minisequencing/primer extension methods.
Physical Characteristics
Ethno-geographic ancestry can be predicted using a set of approximately 70 autosomal SNP markers.04 These ancestry informative markers (AIMs) use proprietary algorithms to partition an individual's AIM-SNP profile into sub-Saharan African, Caucasian, East Asian, and Native American components.04 Y-chromosome SNPs and Alu insertion polymorphisms (that are de facto biallelic UEP markers and can be analyzed by SNP methods) are also promising candidate markers for bio-geographic ancestry prediction.05,06
Eye color, contrary to popular genetics textbook explanations, is a complex trait determined by a number of interacting genes.07 The analysis of SNPs from these interacting genes or regions in close linkage with them can predict the eye color of a DNA donor.07
Degraded DNA
Identification of human remains from the World Trade Center disaster was aided by using a panel of approximately 70 autosomal SNPs whose amplimer lengths were greater than 100 bp and therefore suitable for degraded DNA specimens.08
Additional Online Courses
- What Every First Responding Officer Should Know About DNA Evidence
- Collecting DNA Evidence at Property Crime Scenes
- DNA – A Prosecutor’s Practice Notebook
- Crime Scene and DNA Basics
- Laboratory Safety Programs
- DNA Amplification
- Population Genetics and Statistics
- Non-STR DNA Markers: SNPs, Y-STRs, LCN and mtDNA
- Firearms Examiner Training
- Forensic DNA Education for Law Enforcement Decisionmakers
- What Every Investigator and Evidence Technician Should Know About DNA Evidence
- Principles of Forensic DNA for Officers of the Court
- Law 101: Legal Guide for the Forensic Expert
- Laboratory Orientation and Testing of Body Fluids and Tissues
- DNA Extraction and Quantitation
- STR Data Analysis and Interpretation
- Communication Skills, Report Writing, and Courtroom Testimony
- Español for Law Enforcement
- Amplified DNA Product Separation for Forensic Analysts